RESEARCHIn data-driven computational sciences, a robust image-to-analysis paradigm is critical for the investigation of numerous complex phenomena. The current state-of-the-art in direct image to analysis is constrained by an inability to deal with noisy data, computational inefficiency, data analysis inaccuracy, and limited applications. In the fields of image analysis, geometric modeling, and simulation of real-world phenomena, spline-based techniques are powerful tools for developing smooth and accurate representations of solutions.
|
|
|
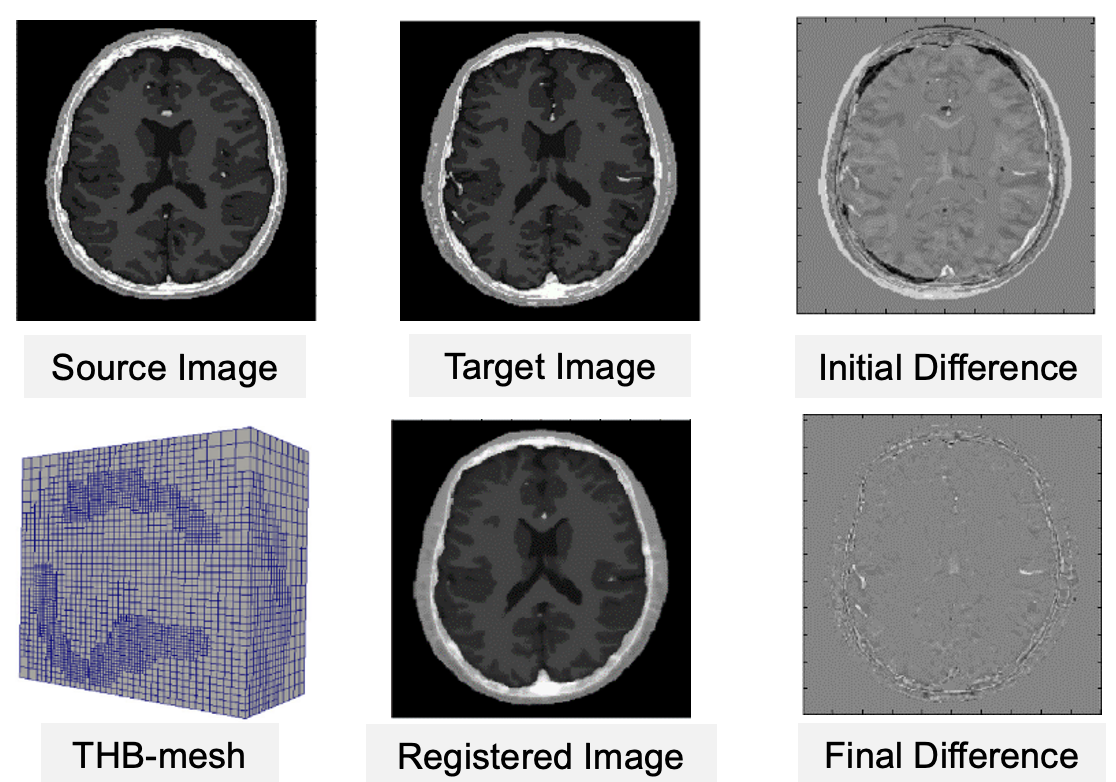
Adaptive Refinement in B-spline Image Registration for Large Deformation Mapping
Clinicians use medical images to study the progression of diseases and provide diagnosis. Due to the large variability within imaging acquisition methods and limitations of image quality, medical image analysis is still a very challenging problem. We are developing novel algorithms for efficiently evaluating large and complex physiological changes between medical images by aligning images acquired at various stages, in order to provide significantly faster and more accurate prediction results, thereby facilitating a better understanding of certain diseases and more personalized treatments. The resulting open-source softwares can be applied to the construction of patient-specific computational models directly from image data. |
|
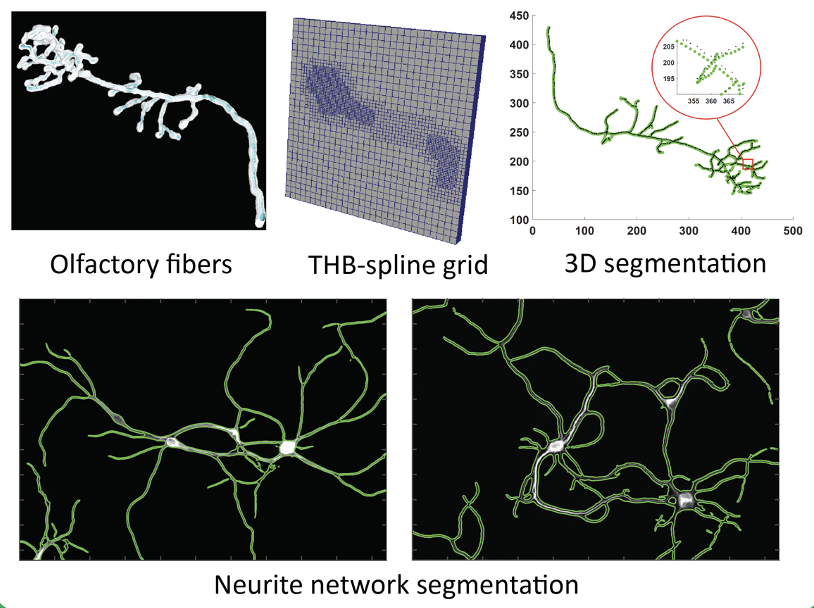
Automatic neuron reconstruction from images
In the field of computational neuroscience, the study of neuron morphology and its growth process is a complex and difficult problem. Understanding the complex morphology development of neurons is critical towards understanding their structural and functional role in the nervous system.We are working on novel frameworks along with open-source softwares for automatic neuron segmentation using a B-spline based active contour deformation model with hyperelastic regularization and automatic initialization. Unlike other existing methods which represent neuron boundary as piecewise constant function, we provide a more accurate and smooth representation of the neuron geometry. |
|
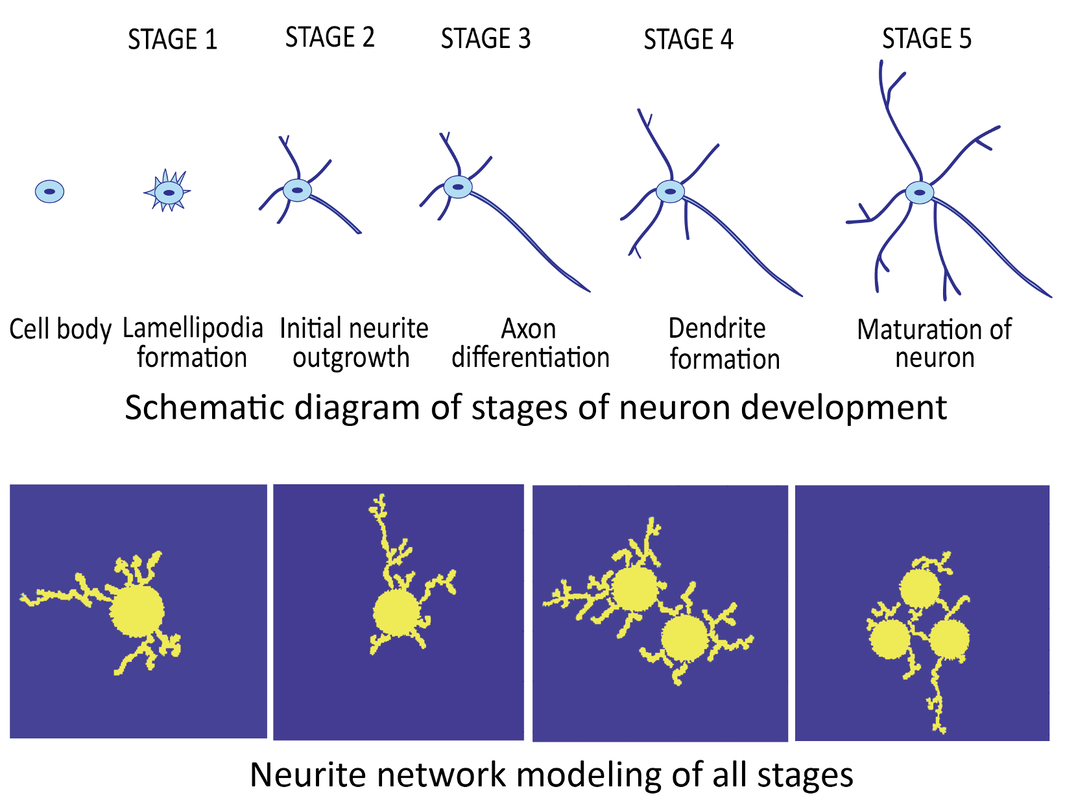
Modeling of Multi-Stage Neuron Morphology Development
We focus on structural modeling of neuron networks and understanding the functional aspects of neuron development, such as the various biochemical and signal pathways governing the development process. We are also looking into the role of mechanics in growth and interactions between neurons. Modeling neuron cell degeneration under the influence of certain factors will enable the research into the study of neurodegenerative diseases such as Alzheimer’s and Parkinson’s, how the abnormalities during development can affect neuron morphology. |
|
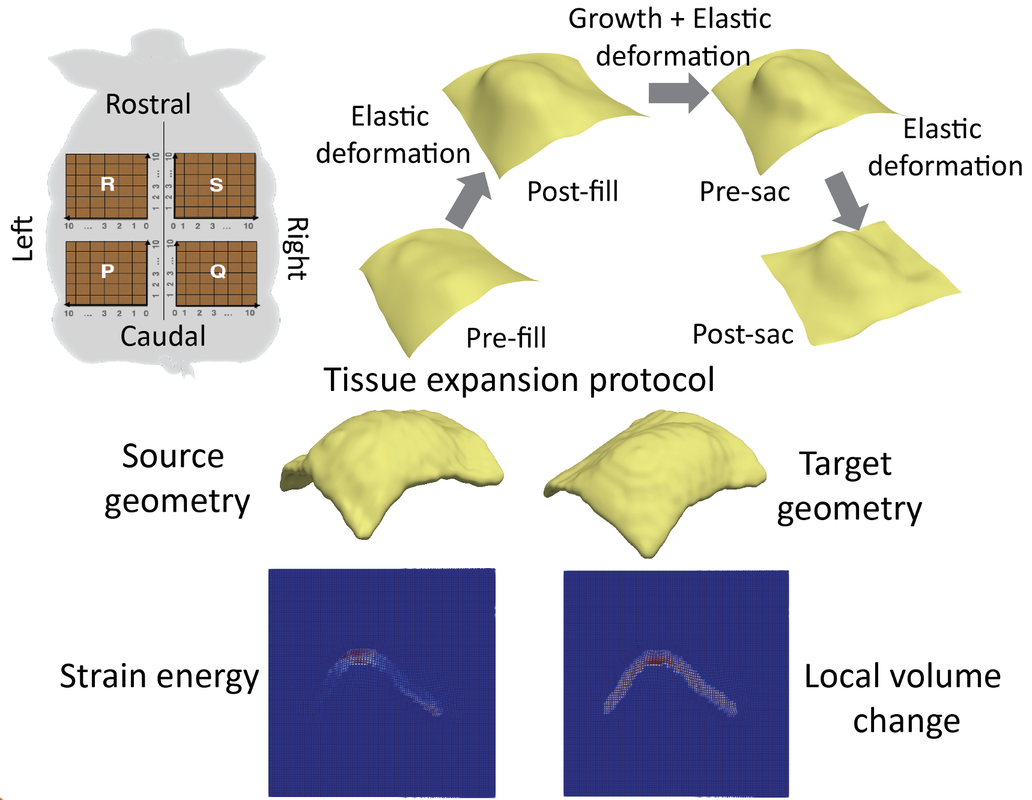
Multiscale Modeling for Biological Tissue Growth, Expansion and Deformation
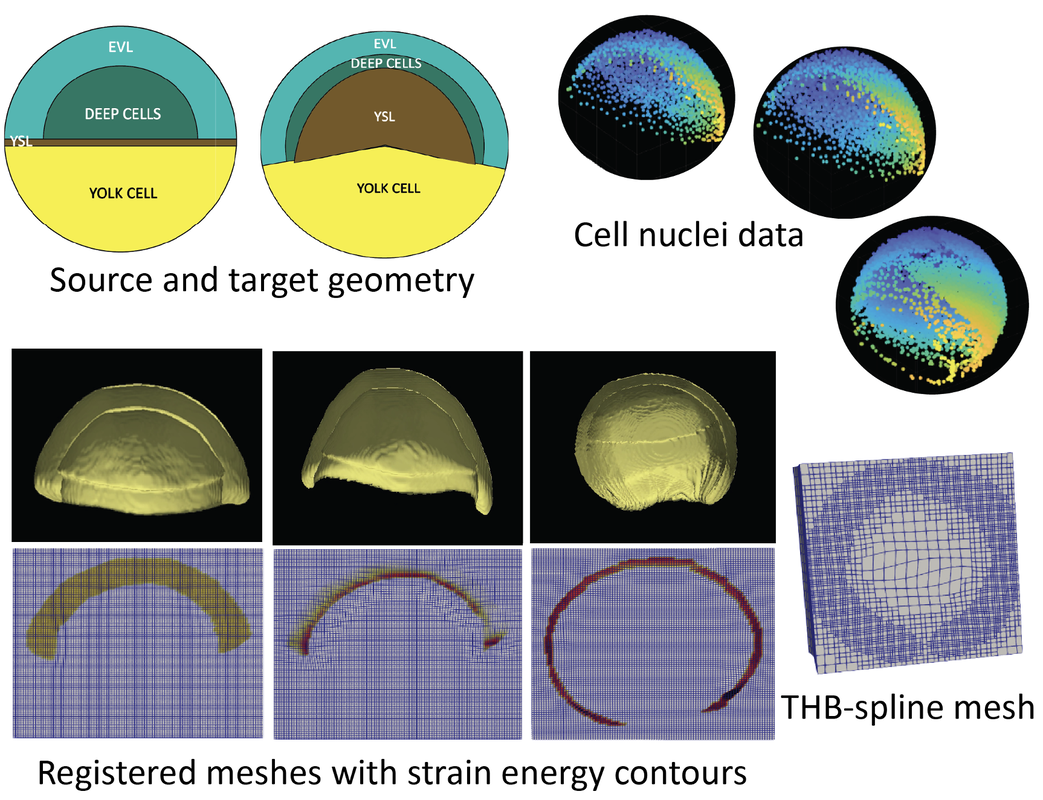
In the study of multiscale modeling of biological tissue growth, applying PDE-constrained shape registration to analyze and predict tissue expansion is a very challenging problem. We have been focusing on integration of biophysical models of growth with shape registration to achieve as realistic as possible tissue deformation. The tissue expansion process is modeled using volume growth theory in continuum mechanics as a PDE constraint. One application is modeling skin growth during tissue expansion reconstructive surgery. Tissue expansion is a common technique for breast reconstruction surgery after mastectomy for breast cancer patients. Computational modeling helps in the prediction of skin growth and adaptation to mechanical cues during the procedure. Through the PDE constrained shape registration framework we can capture complex and large deformation of skin, which is susceptible to anisotropic growth and non-uniform stretching. Another application is computational modeling of epiboly in zebrafish embryos. During embryogenesis, several extracellular factors are responsible for dynamic control and regulation of Bone Morphogenetic Protein (BMPs) signals which are responsible for spatial organization of embryonic axes, organ structure and inducing spatiotemporal patterns of gene expression. The aim is to understand the mechanism of BMP regulation in zebrafish gastrula. Based on the in vivo imaging data, we develop an accurate and optimal spatial mapping that is evaluated between pairs of datasets of cell nuclei captured through different stages of the epiboly process. As a natural extension, this work can be used to study the role of mechanics in different biological processes such as organogenesis, cancer metastasis, angiogenesis and development. |